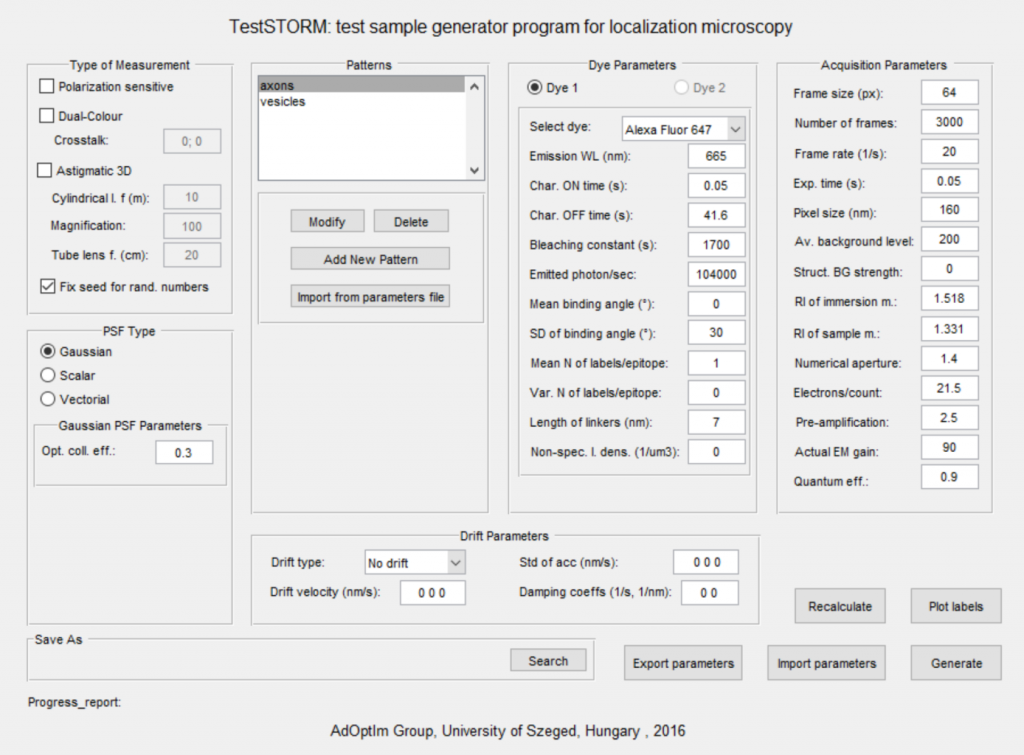
![]() – a simulator written in Mathworks Matlab for modelling the whole imaging procedure in an optical fluorescence localization microscope.
– a simulator written in Mathworks Matlab for modelling the whole imaging procedure in an optical fluorescence localization microscope.
TestSTORM version 2.1.4 (2024. January) – stable
Matlab code: testSTORM-v2.1.5
User Guide: TestSTORM_ver2.1_guide
Release highlights:
v2.1.5 (2024. August)
- Fixed a compatibility issue with newer Matlab versions concerning the vectorial PSF calculation.
v2.1.4 (2024. January)
- Bug fix for for the dual-color simulation.
- Bug fix for the linker orientation in case of the “Disks” pattern.
v2.1.3 (2019. May):
- Performance improvement when running the simulation with large number of labels. Significant memory reduction for the simulations (should be between 2 and 4 times).
- Axial position for the “Array” pattern now can be set in the GUI. It is useful for generating calibration files for the astigmatic 3D simulations.
- FIX: Fixed a bug introduced in the version 2.1 that prevented the reading of input parameters in case of using the Gaussian PSF model and resulted in an error message.
- v2.1.2 Fixed a bug introduced in the version 2.1 that prevented the reading of input parameters in case of using the Gaussian PSF model and resulted in an error message.
- v2.1.2: Fixed a bug affecting the “z” coordinates of the Array pattern. Now the pattern should be symmetric around the z=0 plane.
- v2.1.2: Now the PSF testing tool (“test_TestSTORM_PSF.m”) does not allow polarization sensitive simulation with the Gaussian or with the scalar PSF model.
- v2.1.2: Testing the vectorial PSF model resulted in an error message instead of the pixelized image (“and” condition should have been used instead of the “or” at two place in the code)
- v2.1.2: An error message has been added to hinder starting simulations with inconsistent Exp. time and Frame rate values.
- Replaced Matlab’s iteration to a custom one for increased reliability when calculating scalar or vectorial PSFs.
- Modification to the focal position determination when calculating scalar or vectorial PSFs. Results only slightly different focal position than the previous algorithm, in normal cases (~22 nm difference with the default settings).
- Fixed a bug when calculating scalar or vectorial PSFs (not sure whether a new Matlab version caused it or it was originally in the code). The PSFs shapes and peaks were only marginally affected.
- Made a workaround for a bug causing higher blinking event brightness when simulating with very low “Sample depth” value.
- Added “Synaptosomes” option to the random “Vesicles” pattern.
- Parameters files exported with previous TestSTORM versions (2.0.x) and containing “Rods”, “Vesicles” or “Octagons” patterns can not be imported any longer starting this version.
- Added a tool with which the generated PSF can be depicted.
Other clean-ups and modification in the code which should not affect the behaviour of the program.
TestSTORM version 2.0.4 (2018. June) – old-stable
Matlab code: testSTORM-v2.0.4
User Guide: TestSTORM_ver2.0_guide
Reference: Tibor Novák, Tamás Gajdos, József Sinkó, Gábor Szabó and Miklós Erdélyi: TestSTORM: Versatile simulator software for multimodal super-resolution localization fluorescence microscopy, Sci.Rep., doi: 10.1038/s41598-017-01122-7 (2017)
Release highlights:
- Redesigned GUI.
- New simulated structures: axons, npc (octagons), segmented lines and mesh
- New PSF models: scalar (Gibson and Lani) and vectorial (Richards and Wolf)
- PSF anisotropy
- New simulation types: polarization sensitive, Dual-Colour, Astigmatic 3D
- Structured background
- Drift simulation
- Version controlled release
- v2.0.2: FIX: stl load bugfix
- v2.0.2: New pattern upon user request
- v2.0.3: New disks pattern. (Cylinder shape, with labeled twin-disks inside ~130nm separated.
- v2.0.3: enabled elliptical vesicles in the GUI
- v2.0.3: FIX: random ves. positions, nanorod structure dye orientation, typos,
- v2.0.4: FIX: A serious bug was affecting the PSF size when running simulations subsequently with different parameters (e.g. wavelength).
TestSTORM version 1.0 (2014. Feb.) – obsolete/EOL
Matlab code: testSTORM_ver1_0.zip
Matlab compiled exe: testSTORM_ver1_0pkg.zip
User Guide: TestSTORM_guide.pdf
Reference: Sinkó, J. et. al. TestSTORM: Simulator for optimizing sample labeling and image acquisition in localization based super-resolution microscopy, Biomed. Optics Express 5, 778-787 (2014)
Release highlights:
- Initial release with Gaussian PSF and 4 simulated samples.



You must be logged in to post a comment.